PROVIDENCE, R.I. [Brown University] — It has been known for some time that plants respond to environmental cues that guide their flowering. Chief among these signals are light, temperature and vernalization, when flowering is promoted by prolonged exposure to cold temperatures.
In some plants, scientists have identified particular genes that deal with each of these environmental signals. But they haven't fully grasped how plants integrate these signals in nature. For example, when day length and temperature are combined in different ways, plants outdoors may not respond the same way as plants in the lab.
Through a series of field experiments at five European sites, a Brown University-led research team has charted the internal and external signals that guide the life cycle of one plant species, Arabidopsis thaliana, across its native climate range. The team has created a model that shows the importance of the genetic and environmental cues for key genotypes of Arabidopsis and how these signals vary depending on the plant’s location and seasonal environment.
“This is a powerful tool to predict how this plant species and other species will respond to climate change and which genetic pathways are important in different environments,” said Amity Wilczek, a postdoctoral research associate in ecology and evolutionary biology at Brown and the paper’s lead author.
The paper appears this week in the online edition of Science.
Johanna Schmitt, director of the Environmental Change Initiative at Brown and a professor of biology and environmental studies, is the corresponding author. The contributing authors include Martha Cooper in the Ecology and Evolutionary Biology department and 2006 Brown graduates Laura Martin and Alexis Walker, who were stationed at labs in Europe as part of the project’s international training program for recent graduates interested in pursuing basic research. Other project research fellows listed as authors on the paper include Jillian Anderson, J. Franklin Egan, Cristina Lopez-Gallego, Brook Moyers, Chris Muir, Renee Petipas and Sheina Sim.
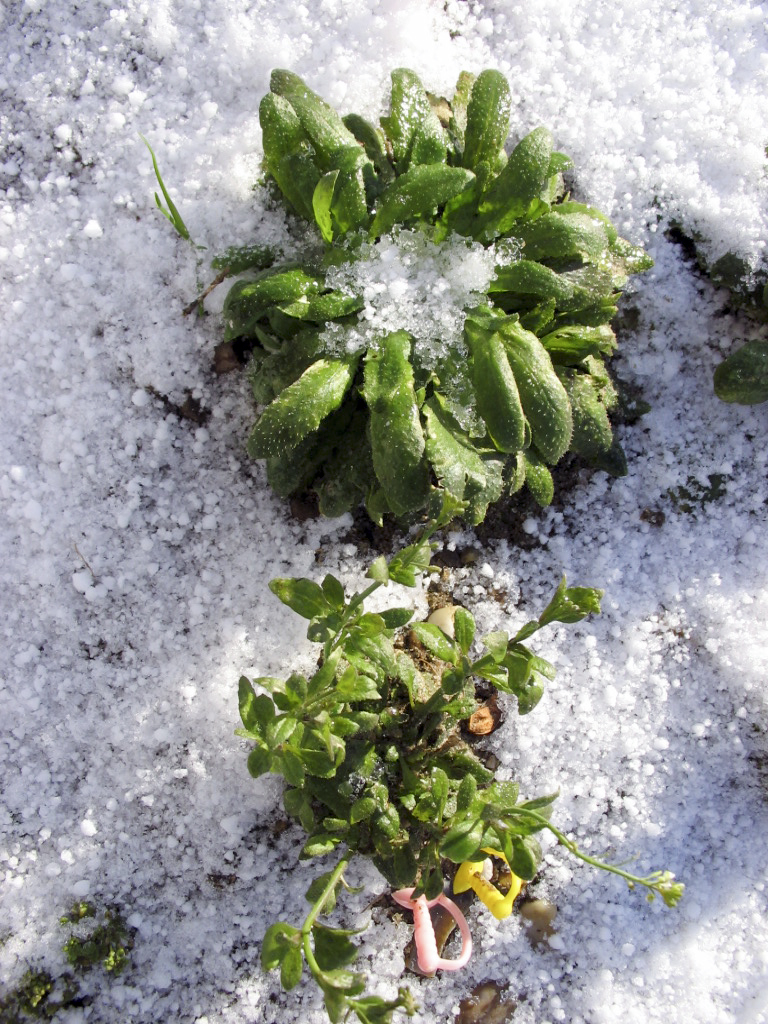
The team used the much-studied plant Arabidopsis, an annual weed closely related to canola and cabbage. The researchers sowed the plant in the wild to study the genetic influences in the plant’s life cycle and how they differed depending on genotype, season and climate. They chose sites spanning the plant’s natural range in Europe: Oulu, Finland; Norwich, United Kingdom; Cologne and Halle, Germany; and Valencia, Spain. They also planted mutants that lacked genes programmed to respond to day length or temperature. The group charted each genotype’s rate of development from planting to bolting (when it begins to flower) and compared results. The experiments spanned several growing seasons, from summer 2006 to fall 2008.
A plant’s internal genetic timer is complex. There is a set of genes programmed to respond to environmental cues such as long days, but there is also another class of repressor genes that delay or stop the environmentally cued genes from working. This seesaw competition of genetic pathways inside the plant goes on until a threshold is reached, a molecular switch that triggers the plant to cross into the next stage of development. By examining mutants impaired in different genes, the team could quantify shifts in the balance of this seesaw across different seasons and climates. The team discovered that certain mutations with major effects under laboratory conditions had variable and sometimes unexpectedly small effects in natural field environments.

Wilczek and Schmitt, along with Stephen Welch, a professor in the agronomy department at Kansas State University, then created models that charted precisely the rates of development for the genotypes at the field sites, using hourly temperature and light data collected at each site. They showed when each genotype would reach its threshold and switch from a vegetative state to a flowering stage. The models also accurately predicted the contribution of each genetic pathway to development and how the pathways are affected by environmental cues.
“With our model, we have shown that we can successfully predict how flowering is going to behave under a range of environmental conditions, not just those in which we originally grew our plants,” Wilczek said. “Given the changing climate and the importance of flowering timing for wild plants and crop plants, this model can help us better understand how plants will respond to future conditions.”
What became apparent, Wilczek added, is that “both genes and environments are important to figure out what plants are going to do. You really need both.”
Schmitt said the modeling approach could be used for other plant species, including crops and ecologically important wild species. She also believes the model can be tweaked to predict how other plants may respond to changing climate. “There are a lot of possible parallels,” Schmitt said.
“Future climates will have different combinations of temperature and light periods,” Wilczek added, “so it‘s good to have models to deal with those so-called non-analog climates.”
Other contributing authors included Judith Roe, Mary Knapp and Erika Charbit from Kansas State University, and George Coupland and Antonis Giakountis of the Max Planck Institute for Plant Breeding Research in Cologne, Germany.
The National Science Foundation and the Alexander von Humboldt Foundation funded the research.